What is Exploratory Data Analysis (EDA) and How Does it Work?

Introduction
Exploratory Data Analysis (EDA) is a process of describing the data by means of statistical and visualization techniques in order to bring important aspects of that data into focus for further analysis. This involves inspecting the dataset from many angles, describing & summarizing it without making any assumptions about its contents.
“Exploratory Data analysis is an attitude, a state of flexibility, a willingness to look for those things that we believe are not there, as well as those we believe to be there” – John W. Tukey
EDA is a significant step to take before diving into statistical modeling or machine learning, to ensure the data is really what it is claimed to be and that there are no obvious errors. It should be part of data science projects in every organization.
This article was published as a part of the Data Science Blogathon
Table of contents
- Introduction
- What is Exploratory Data Analysis?
- Types of EDA
- What is Exploratory data analysis Python?
- Why Exploratory Data Analysis is important?
- Data set description
- 1. Importing libraries and loading data
- 2. Understanding data
- 3. Uni-variate data analysis
- 4. Bi-variate data analysis
- 5. Multivariate analysis with Contour plot
- Conclusion
- Frequently Asked Questions
What is Exploratory Data Analysis?
Exploratory Data Analysis (EDA) is like exploring a new place. You look around, observe things, and try to understand what’s going on. Similarly, in EDA, you look at a dataset, check out the different parts, and try to figure out what’s happening in the data.
Here’s what you typically do:
- Look at the Data: You start by gathering information about the data you have. How many rows and columns are there? What kind of information does each column contain?
- Clean the Data: Sometimes, data can be messy. There might be missing values, or some values might be wrong. You clean up the data by fixing these issues.
- Make Summaries: You summarize the data to get a general idea of what’s in it. You might find out things like the average value, the most common value, or how the values are spread out.
- Visualize the Data: It’s helpful to see the data in graphs or charts. This way, you can spot trends or patterns more easily. Python has libraries like Pandas, NumPy, and Matplotlib that are commonly used for this purpose in exploratory data analysis Python.
- Ask Questions: As you explore the data, you might come up with questions. Why is one part of the data different from the rest? Are there any relationships between different parts of the data?
- Find Answers: You try to answer the questions you’ve asked by digging deeper into the data. This might involve doing more analysis or creating models.
Types of EDA
Here are five types of EDA techniques:
- Univariate Analysis: In EDA Analysis, univariate analysis examines individual variables to understand their distributions and summary statistics.
- Bivariate Analysis: This aspect of EDA explores the relationship between two variables, uncovering patterns through techniques like scatter plots and correlation analysis.
- Visualization Techniques: EDA relies heavily on visualization methods to depict data distributions, trends, and associations using various charts and graphs.
- Outlier Detection: EDA involves identifying outliers within the data, anomalies that deviate significantly from the rest, employing tools such as box plots and z-score analysis.
- Statistical Tests: EDA often includes performing statistical tests to validate hypotheses or discern significant differences between groups, adding depth to the analysis process.
What is Exploratory data analysis Python?
Exploratory Data Analysis (EDA) is an essential step in the data analysis process. It involves analyzing and visualizing data to understand its main characteristics, uncover patterns, and identify relationships between variables. Python offers several libraries that are commonly used for EDA, including pandas, NumPy, Matplotlib, Seaborn, and Plotly. Here’s a basic example of how you can perform EDA using Python:
# Import necessary libraries
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
# Load the dataset
data = pd.read_csv('your_dataset.csv')
# Display basic information about the dataset
print("Shape of the dataset:", data.shape)
print("\nColumns:", data.columns)
print("\nData types of columns:\n", data.dtypes)
# Display summary statistics
print("\nSummary statistics:\n", data.describe())
# Check for missing values
print("\nMissing values:\n", data.isnull().sum())
# Visualize distribution of a numerical variable
plt.figure(figsize=(10, 6))
sns.histplot(data['numerical_column'], kde=True)
plt.title('Distribution of Numerical Column')
plt.xlabel('Numerical Column')
plt.ylabel('Frequency')
plt.show()
# Visualize relationship between two numerical variables
plt.figure(figsize=(10, 6))
sns.scatterplot(x='numerical_column_1', y='numerical_column_2', data=data)
plt.title('Relationship between Numerical Column 1 and Numerical Column 2')
plt.xlabel('Numerical Column 1')
plt.ylabel('Numerical Column 2')
plt.show()
# Visualize relationship between a categorical and numerical variable
plt.figure(figsize=(10, 6))
sns.boxplot(x='categorical_column', y='numerical_column', data=data)
plt.title('Relationship between Categorical Column and Numerical Column')
plt.xlabel('Categorical Column')
plt.ylabel('Numerical Column')
plt.show()
# Visualize correlation matrix
plt.figure(figsize=(10, 6))
sns.heatmap(data.corr(), annot=True, cmap='coolwarm', fmt=".2f")
plt.title('Correlation Matrix')
plt.show()
Why Exploratory Data Analysis is important?
Just like everything in this world, data has its imperfections. Raw data is usually skewed, may have outliers, or too many missing values. A model built on such data results in sub-optimal performance. In hurry to get to the machine learning stage, some data professionals either entirely skip the exploratory data analysis (EDA) process or do a very mediocre job. This is a mistake with many implications, that includes generating inaccurate models, generating accurate models but on the wrong data, not creating the right types of variables in data preparation, and using resources inefficiently.
In this article, we’ll be using Pandas, Seaborn, and Matplotlib libraries of Python to demonstrate various EDA techniques applied to Haberman’s Breast Cancer Survival Dataset.
Data set description
The dataset comprises cases from the exploratory data analysis conducted between 1958 and 1970 at the University of Chicago’s Billings Hospital, focusing on the survival of patients post-surgery for breast cancer
The dataset can be downloaded from here. [Source: Tjen-Sien Lim ([email protected]), Date: March 4, 1999]
Attribute information :
- Patient’s age at the time of operation (numerical).
- Year of operation (year — 1900, numerical).
- A number of positive axillary nodes were detected (numerical).
- Survival status (class attribute)
1: the patient survived 5 years or longer post-operation.
2: the patient died within 5 years post-operation.
Attributes 1, 2, and 3 form our features (independent variables), while attribute 4 is our class label (dependent variable).
Let’s begin our analysis . . .
1. Importing libraries and loading data
Import all necessary packages —
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import scipy.stats as statsLoad the dataset in pandas dataframe —
df = pd.read_csv('haberman.csv', header = 0)
df.columns = ['patient_age', 'operation_year', 'positive_axillary_nodes', 'survival_status']2. Understanding data
Output:
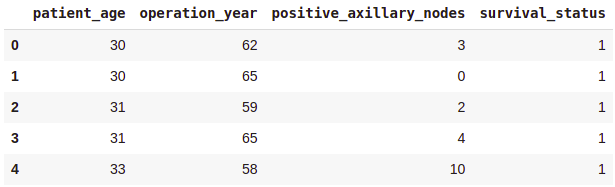
Shape of the dataframe —
df.shapeOutput:
(305, 4)
There are 305 rows and 4 columns. But how many data points for each class label are present in our dataset?
df[‘survival_status’].value_counts()Output:

- The dataset is imbalanced as expected.
- Out of a total of 305 patients, the number of patients who survived over 5 years post-operation is nearly 3 times the number of patients who died within 5 years.
df.info()
Output:
- All the columns are of integer type.
- No missing values in the dataset.
2.1 Data preparation
Before we go for statistical analysis and visualization, we see that the original class labels — 1 (survived 5 years and above) and 2 (died within 5 years) are not in accordance with the case.
So, we map survival status values 1 and 2 in the column survival_status to categorical variables ‘yes’ and ‘no’ respectively such that,
survival_status = 1 → survival_status = ‘yes’
survival_status = 2 → survival_status = ‘no’
df['survival_status'] = df['survival_status'].map({1:"yes", 2:"no"})2.2 General statistical analysis
df.describe()Output:
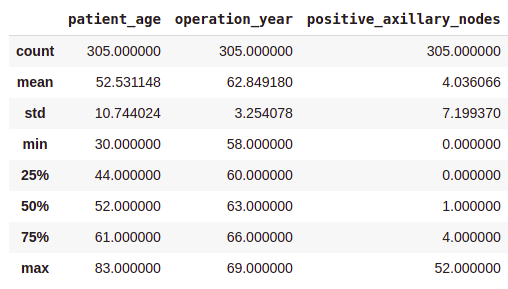
- On average, patients got operated at age of 63.
- An average number of positive axillary nodes detected = 4.
- As indicated by the 50th percentile, the median of positive axillary nodes is 1.
- As indicated by the 75th percentile, 75% of the patients have less than 4 nodes detected.
If you see, there is a significant difference between the mean and the median values. This is because there are some outliers in our data and the mean is influenced by the presence of outliers.
2.3 Class-wise statistical analysis
survival_yes = df[df['survival_status'] == 'yes']
survival_yes.describe()Output:
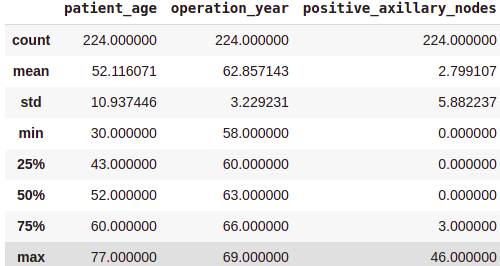
survival_no = df[df[‘survival_status’] == ‘no’] survival_no.describe()
Output:
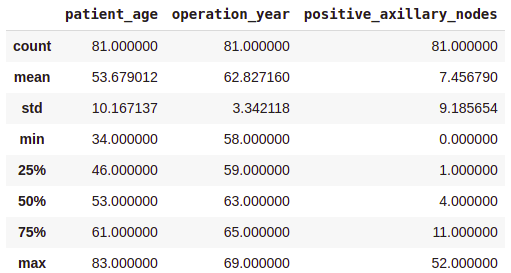
From the above class-wise analysis, it can be observed that —
- The average age at which the patient is operated on is nearly the same in both cases.
- Patients who died within 5 years on average had about 4 to 5 positive axillary nodes more than the patients who lived over 5 years post-operation.
Note that, all these observations are solely based on the data at hand.
3. Uni-variate data analysis
“A picture is worth ten thousand words”
– Frank R. Bernard
3.1 Distribution Plots
Uni-variate analysis, as the name suggests, involves studying one variable at a time. Suppose our objective is to accurately ascertain the survival status based on features such as patient’s age, operation year, and positive axillary nodes count. In this context of EDA (exploratory data analysis), determining the most informative variable among these three becomes crucial for distinguishing between the class labels ‘yes’ and ‘no.’ To address this question, we’ll create distribution plots, also known as probability density function (PDF) plots, with each feature representing a variable on the X-axis. The values on the Y-axis in each case depict the normalized density.
Patients’s age
sns.FacetGrid(df, hue = "survival_status").map(sns.distplot, "patient_age").add_legend() plt.show()Output:
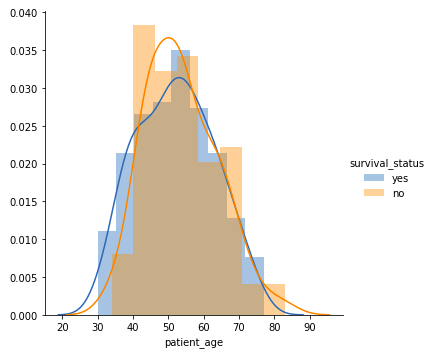
- Among all the age groups, the patients belonging to 40-60 years of age are the highest.
- There is a high overlap between the class labels. This implies that the survival status of the patient post-operation cannot be discerned from the patient’s age.
Operation year
FacetGrid(df, hue = "survival_status").map(sns.distplot, "operation_year").add_legend() plt.show()Output:
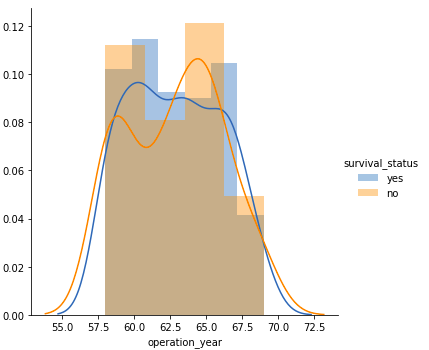
Just like the above plot, here too, there is a huge overlap between the class labels suggesting that one cannot make any distinctive conclusion regarding the survival status based solely on the operation year.
Number of positive axillary nods.
FacetGrid(df, hue = "survival_status").map(sns.distplot, "positive_axillary_nodes").add_legend() plt.show()Output:
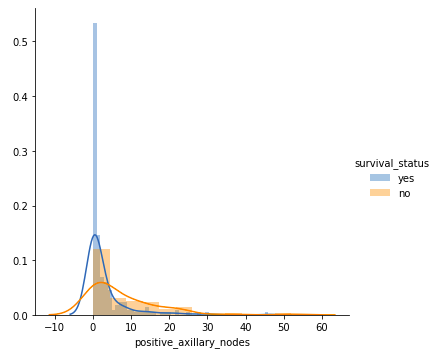
This plot looks interesting! Although there is a good amount of overlap, here we can make some distinctive observations –
- Patients having 4 or fewer axillary nodes — A very good majority of these patients have survived 5 years or longer.
- Patients having more than 4 axillary nodes — the likelihood of survival is found to be less as compared to the patients having 4 or fewer axillary nodes.
But our observations must be backed by some quantitative measure. That’s where the Cumulative Distribution function(CDF) plots come into the picture.
The area under the plot of PDF over an interval represents the probability of occurrence of the random variable in the given interval. Mathematically, CDF is an integral of PDF over the range of values that a continuous random variable takes. CDF of a random variable at any point ‘x’ gives the probability that a random variable will take a value less than or equal to ‘x’.
counts, bin_edges = np.histogram(survival_yes[‘positive_axillary_nodes’], density = True) pdf = counts/sum(counts) cdf = np.cumsum(pdf) plt.plot(bin_edges[1:], cdf, label = ‘CDF Survival status = Yes’)
counts, bin_edges = np.histogram(survival_no[‘positive_axillary_nodes’], density = True) pdf = counts/sum(counts) cdf = np.cumsum(pdf) plt.plot(bin_edges[1:], cdf, label = ‘CDF Survival status = No’) plt.legend() plt.xlabel(“positive_axillary_nodes”) plt.grid() plt.show()
Output:
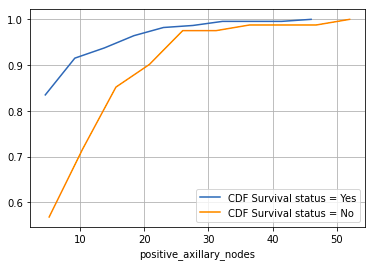
Some of the observations that could be made from the CDF plot —
- Patients having 4 or fewer positive axillary nodes have about 85% chance of survival for 5 years or longer post-operation, whereas this number is less for the patients having more than 4 positive axillary nodes. This gap diminishes as the number of axillary nodes increases.
3.2 Box plots and Violin plots
The box plot, commonly referred to as a box and whisker plot, serves as a visual representation that summarizes exploratory data analysis Python using five key metrics — the minimum, lower quartile (25th percentile), median (50th percentile), upper quartile (75th percentile), and maximum data values.
A violin plot displays the same information as the box and whisker plot; additionally, it also shows the density-smoothed plot of the underlying distribution.
Let’s make the box plots for our feature variables
plt.figure(figsize = (15, 4)) plt.subplot(1,3,1) sns.boxplot(x = ‘survival_status’, y = ‘patient_age’, data = df) plt.subplot(1,3,2) sns.boxplot(x = ‘survival_status’, y = ‘operation_year’, data = df) plt.subplot(1,3,3) sns.boxplot(x = ‘survival_status’, y = ‘positive_axillary_nodes’, data = df) plt.show()
Output:
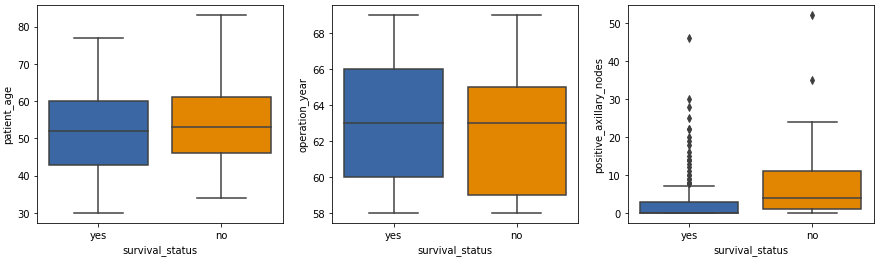
- The patient age and the operation year plots show similar statistics.
- The isolated points seen in the box plot of positive axillary nodes are the outliers in the data. Such a high number of outliers is kind of expected in medical datasets.
Violin Plots -plt.figure(figsize = (15, 4)) plt.subplot(1,3,1) sns.violinplot(x = ‘survival_status’, y = ‘patient_age’, data = df) plt.subplot(1,3,2) sns.violinplot(x = ‘survival_status’, y = ‘operation_year’, data = df) plt.subplot(1,3,3) sns.violinplot(x = ‘survival_status’, y = ‘positive_axillary_nodes’, data = df) plt.show()
Output:
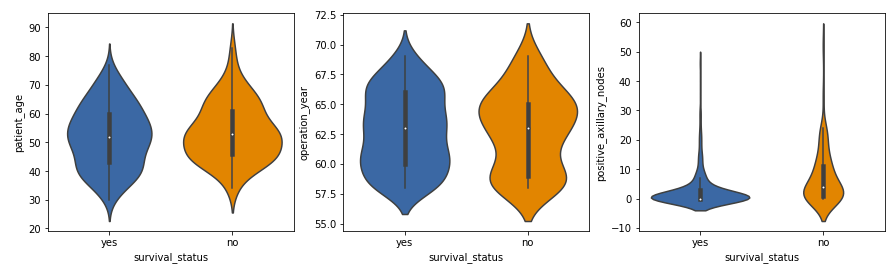
Violin plots
A powerful tool in exploratory data analysis, offer greater insights compared to traditional box plots. These plots not only present a statistical summary but also visually depict the underlying distribution of the data. Examining the violin plot for positive axillary nodes, it becomes apparent that the distribution is highly skewed for the ‘yes’ class label and moderately skewed for the ‘no’ label. This indicates that –
- For the majority of patients (in both the classes), the number of positive axillary nodes detected is on the lesser side. Of which, patients having 4 or fewer positive axillary nodes are more likely to survive 5 years post-operation.
These observations are consistent with our observations from previous sections.
4. Bi-variate data analysis
4.1 Pair plot
Next, we shall plot a pair plot to visualize the relationship between the features in a pairwise manner. A pair plot enables us to visualize both distributions of single variables as well as the relationship between pairs of variables.sns.set_style(‘whitegrid’) sns.pairplot(df, hue = ‘survival_status’) plt.show()
Output:
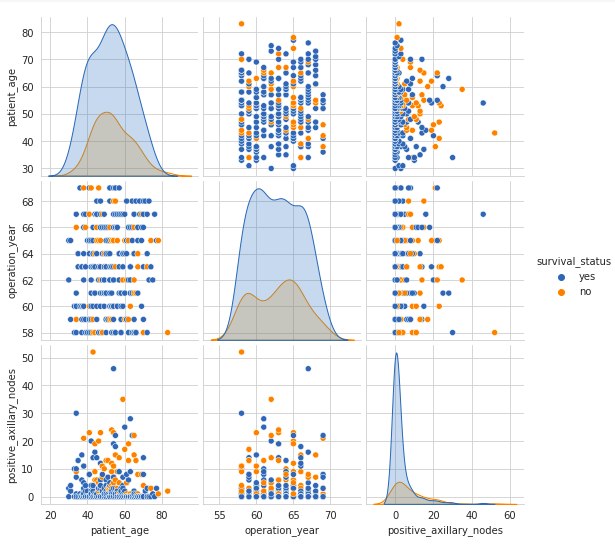
In the context of exploratory data analysis, examining the pair plot reveals that the upper half and lower half plots on the diagonal interchange axes but convey identical information. Therefore, analyzing either set is sufficient for gaining insights. Notably, the plots on the diagonal differ from the others, showcasing kernel density smoothed histograms that depict the univariate distribution of specific features
As we can observe in the above pair plot, there is a high overlap between any two features and hence no clear distinction can be made between the class labels based on the feature pairs.
4.2 Joint plot
While the Pair plot provides a visual insight into all possible correlations, the Joint plot provides bivariate plots with univariate marginal distributions.sns.jointplot(x = ‘patient_age’, y = ‘positive_axillary_nodes’, data = df) plt.show()
Output:
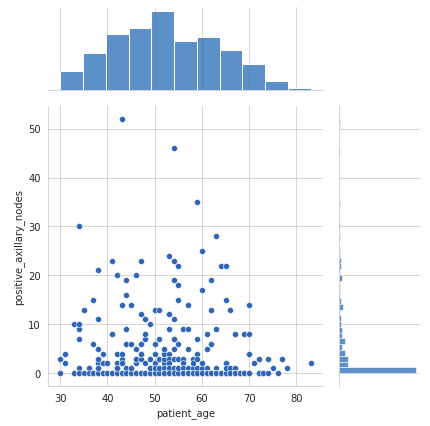
- The pair plot and the joint plot reveal that there is no correlation between the patient’s age and the number of positive axillary nodes detected.
- The histogram on the top edge indicates that patients are more likely to get operated in the age of 40–60 years compared to other age groups.
- The histogram on the right edge indicates that the majority of patients had fewer than 4 positive axillary nodes.
4.3 Heatmap
Heatmaps play a crucial role in exploratory data analysis, allowing us to visually assess correlations among feature variables. EDA data analysis becomes especially significant when aiming to determine the feature importance in regression analysis. While EDA correlated features might not directly affect the statistical model’s performance, they can introduce complexity into the post-modeling analysis
Let’s see if there exist any correlation among our features by plotting a heatmap.
sns.heatmap(df.corr(), cmap = ‘YlGnBu’, annot = True)
plt.show()
Output:
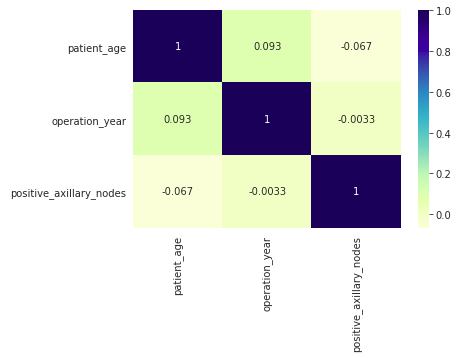
The values in the cells are Pearson’s R values which indicate the correlation among the feature variables. As we can see, eda data analysis these values are nearly 0 for any pair, so no correlation exists among any pair of variables.
5. Multivariate analysis with Contour plot
A contour plot, a valuable tool in exploratory data analysis, is a graphical technique for representing a 3-dimensional surface by plotting constant z slices, called contours, in a 2-dimensional format. This visualization method allows us to efficiently consolidate information from the 3rd dimension into a flat 2-D chart –
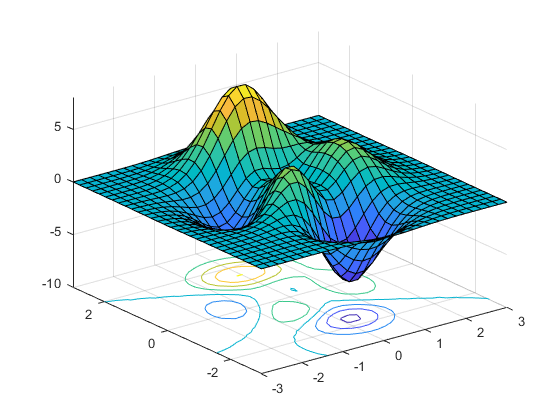
1. Patient’s age
Contour plot example | Source: https://www.mathworks.com/help/matlab/ref/surfc.html
Plotting a contour plot using the seaborn library for patient’s age on x-axis and operation year on the y-axis —
sns.jointplot(x = 'patient_age', y = 'operation_year' , data = df, kind = 'kde', fill = True)
plt.show()Output:
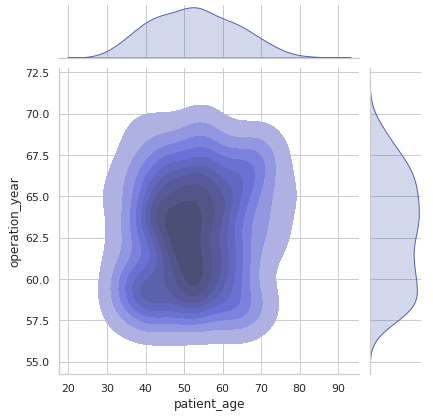
From the above contour plot, it can be observed that the years 1959–1964 witnessed more patients in the age group of 45–55 years.
Conclusion
In this article, we learned some common steps involved in exploratory data analysis. We also saw several types of charts & plots and what information is conveyed by each of these. This is just not it, I encourage you to play with the data and come up with different kinds of visualizations and observe what insights you can extract from it.
Frequently Asked Questions
A. An example of exploratory data analysis (EDA) could involve examining a dataset of customer demographics and purchase history for a retail business. EDA techniques may include calculating summary statistics, visualizing data distributions, identifying outliers, exploring relationships between variables, and performing hypothesis testing. This process helps gain insights into the data, identify patterns, and inform further analysis or decision-making.
A. The four steps of exploratory data analysis (EDA) typically involve:
1. Data Cleaning: Handling missing values, removing outliers, and ensuring data quality.
2. Data Exploration: Examining summary statistics, visualizing data distributions, and identifying patterns or relationships.
3. Feature Engineering: Transforming variables, creating new features, or selecting relevant variables for analysis.
4. Data Visualization: Presenting insights through plots, charts, and graphs to communicate findings effectively.
1. Summarize the Data: Calculate basic statistics for numerical variables and determine frequency distribution for categorical variables.
2. Visualize the Data: Create histograms, scatter plots, box plots, bar charts, and pie charts.
3. Identify Outliers: Detect and investigate outliers using statistical methods or visualization techniques.
4. Transform the Data: Apply transformations to improve the performance of machine learning algorithms and handle missing values.
5. Identify Relationships: Calculate correlation coefficients and create correlation matrices.
6.Generate Hypotheses: Formulate hypotheses about the underlying patterns and relationships in the data.
7. Iterate and Refine: EDA is an iterative process, so revisit previous steps and refine your analysis as needed.
Data Cleaning: Address missing values and outliers for reliable analysis.
Bivariate Analysis: Explore relationships between key variables for insights.
Data Visualization: Create visuals to understand and communicate patterns.
Feature Engineering: Transform variables to enhance predictive power.
Hypothesis Testing: Formulate and test hypotheses to draw meaningful conclusions.









